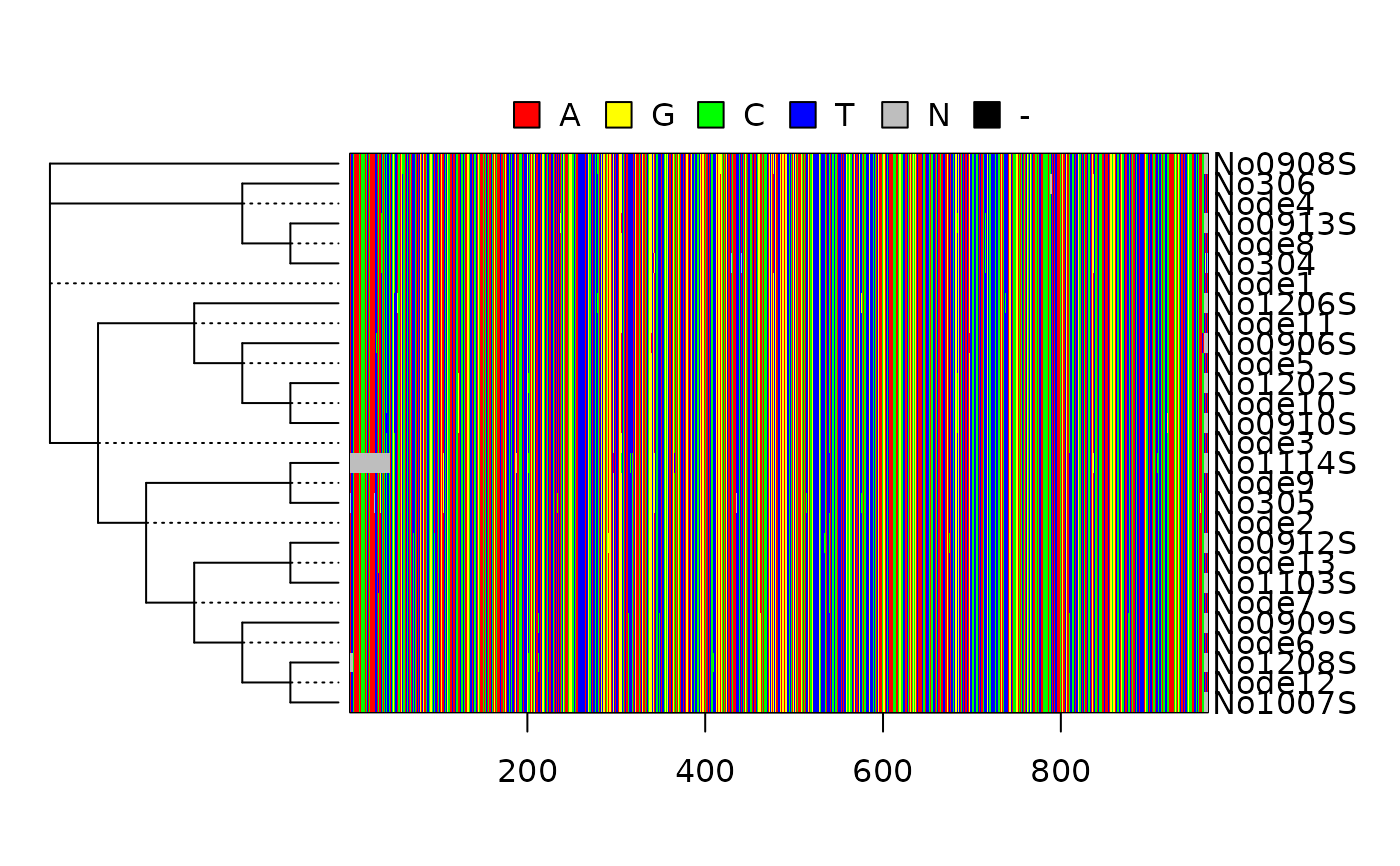
Plots a heat map with ancestral states.
Usage
anc_heatmap(x, y = NULL, use.edge.length = FALSE, align_label = TRUE,
clade = NULL, select = NULL, cex.lab = 1, ...)Arguments
- x
an object of class ancestral or phyDat.
- y
an object of class phyDat.
- use.edge.length
a logical indicating whether to use the edge lengths of the phylogeny to draw the branches (the default) or not (if FALSE). This option has no effect if the object of class "phylo" has no ‘edge.length’ element.
- align_label
a logical value or an integer. If TRUE, the tips are aligned and dotted lines are drawn between the nodes of the tree and the labels.
- clade
a node number or label to extract the clade from the tree.
- select
a subset of characters, columns in the alignment.
- cex.lab
font size of the labels.
- ...
Further arguments passed to or from other methods.
Author
Klaus Schliep klaus.schliep@gmail.com